Adding shared environment (C)¶
The Minimal ACE baseline had \(C = 0\) in both traits. This page configures a single simulation containing two traits with identical \(A\) but contrasting shared-environment variance: trait 1 has \(C = 0\) and trait 2 has \(C = 0.2\). The effect of the C component is then read off the within-scenario trait 1 ↔ trait 2 comparison in the atlas plots.
Configuration¶
with_c:
seed: 90002
replicates: 1
pedigree:
trait1:
A: 0.5
C: 0.0
E: 0.5
trait2:
A: 0.5
C: 0.2
E: 0.3
Both traits have \(A = 0.5\) and total variance \(1.0\), so the input heritability is \(h^2 = 0.5\) in both. Trait 2 reassigns \(0.2\) units of variance from \(E\) to \(C\); cross-trait correlations \(r_A\), \(r_C\), \(r_E\) default to zero, so the two traits are statistically independent.
Run¶
Comparison between trait 1 and trait 2¶
The relative-pair correlation page from
results/examples/with_c/plots/atlas.pdf:
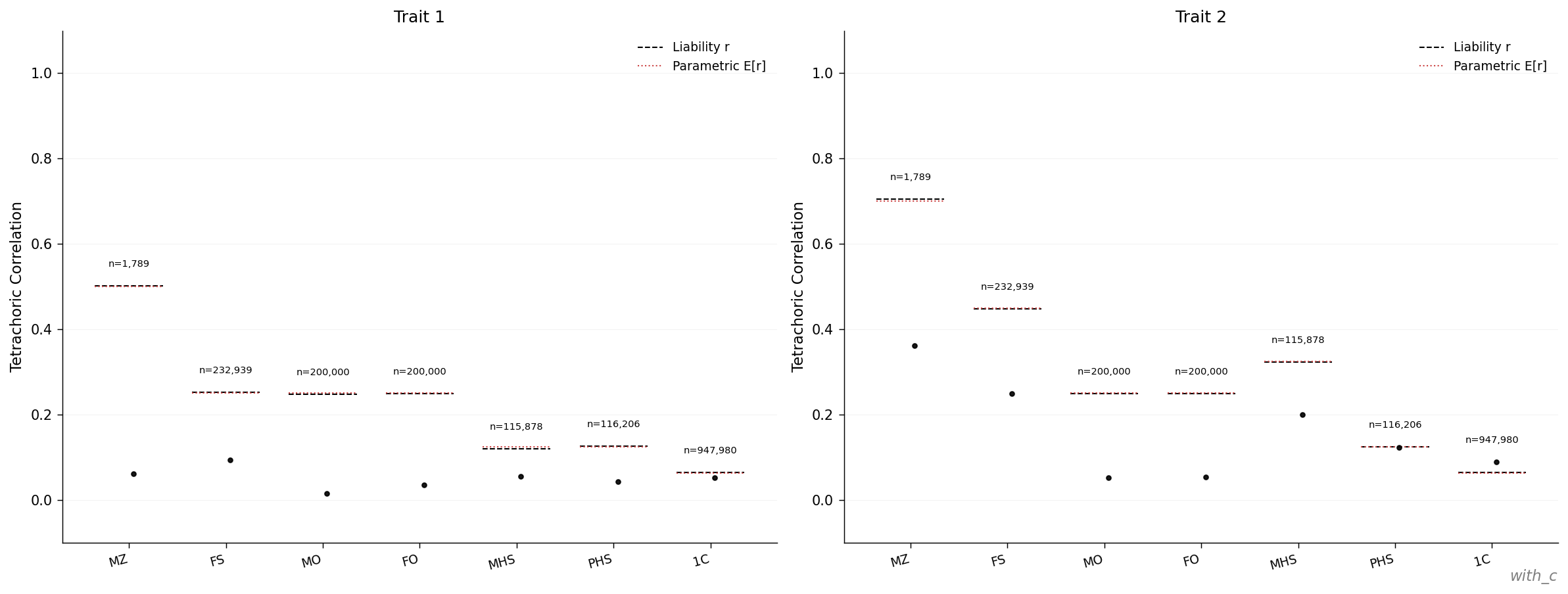
In simACE the C component is shared per household, where a household
is defined by the mother (simace/simulation/simulate.py:761). Full
siblings and maternal half-siblings therefore share C, whereas
parents and their offspring, paternal half-siblings, and cousins draw
C independently.
| Class | Trait 1 (C=0) | Trait 2 (C=0.2) | Trait 2 expectation | Shares C? |
|---|---|---|---|---|
| MZ | 0.50 | 0.70 | \(v_A + v_C = 0.70\) | yes |
| FS | 0.25 | 0.45 | \(0.5\,v_A + v_C = 0.45\) | yes (same mother) |
| MO | 0.25 | 0.25 | \(0.5\,v_A = 0.25\) | no (different generation) |
| FO | 0.25 | 0.25 | \(0.5\,v_A = 0.25\) | no |
| MHS | 0.125 | 0.325 | \(0.25\,v_A + v_C = 0.325\) | yes (same mother) |
| PHS | 0.125 | 0.125 | \(0.25\,v_A = 0.125\) | no (different mothers) |
| 1C | 0.0625 | 0.0625 | \(0.0625\,v_A = 0.0625\) | no |
The variance-component checks in validation.yaml recover \((0.5, 0.0,
0.5)\) for trait 1 and \((0.5, 0.2, 0.3)\) for trait 2, in agreement with
the configured inputs.
Observations¶
Observation 1 — C inflates only the relative classes that share a mother¶
MZ, FS, and MHS each gain exactly \(v_C = 0.2\) between trait 1 and trait 2; MO, FO, PHS, and 1C are unchanged. The increment is the same in every class that shares C, regardless of the additional contribution from the additive component, because the C component is shared in full or not at all.
Observation 2 — The MZ–FS gap is invariant to \(v_C\)¶
The difference \(r_{MZ} - r_{FS} = 0.5 \, v_A\) is independent of \(v_C\), because both MZ and FS share C in full and the \(v_C\) term cancels. Consequently, Falconer's formula \(2(r_{MZ} - r_{FS})\) estimates \(h^2\) rather than \(h^2 + c^2\), and both traits return \(\approx 0.5\) from Falconer despite their materially different relative-pair correlation patterns.
Observation 3 — The MHS–PHS contrast is a direct estimator of \(v_C\)¶
Maternal and paternal half-siblings share the same \(0.25 \, v_A\) by relatedness; the only structural difference between them in this model is that MHS share C and PHS do not. It follows that \(r_{MHS} - r_{PHS} = v_C\) exactly. The trait 2 panel gives \(0.325 - 0.125 = 0.20\), matching the configured input; the trait 1 panel gives \(0\), also as expected. This contrast is the identifiability condition that allows the ACE model to separate \(v_C\) from \(v_E\) (see the ACE Model concept page), and it relies on the maternal household structure used by simACE.
The case in which the clean C decomposition breaks — for example, under assortative mating, which moves variance into a covariance between mates — is treated in AM and Heritability.